14 Conventions and Style
Why names (and style) are part of your statistical method
14.1 Why you should care (even if you “just want to model stuff”)
Naming conventions and coding style are not decoration. They are infrastructure.
If your names are inconsistent, your code becomes harder to read, harder to debug, and harder to share—especially once your project grows beyond “one file, one afternoon.”
“It’s the hardest job to come up with new and unique names for a variable every time you create one but this is the difference between an average programmer and a good one.”
— Vikram Singh Rawat
In scientific workflows, this difference is not about elegance. It is about traceability.
A workflow that depends on unstable or unclear names cannot be audited, reused, or trusted.
Before we talk about modeling or automation, we need to ensure that the basic objects in our workflow—variables, files, and folders—are named in a way that makes their roles unambiguous.
This is what allows:
- metadata to point to the correct data,
- scripts to operate predictably,
- and workflows to declare dependencies explicitly.
14.2 Naming conventions for variables and files
14.2.1 What are “naming conventions”?
They are simply a consistent set of rules for naming:
- variables (columns in your dataset)
- objects in R (data frames, models, plots)
- files and folders (scripts, data, outputs)
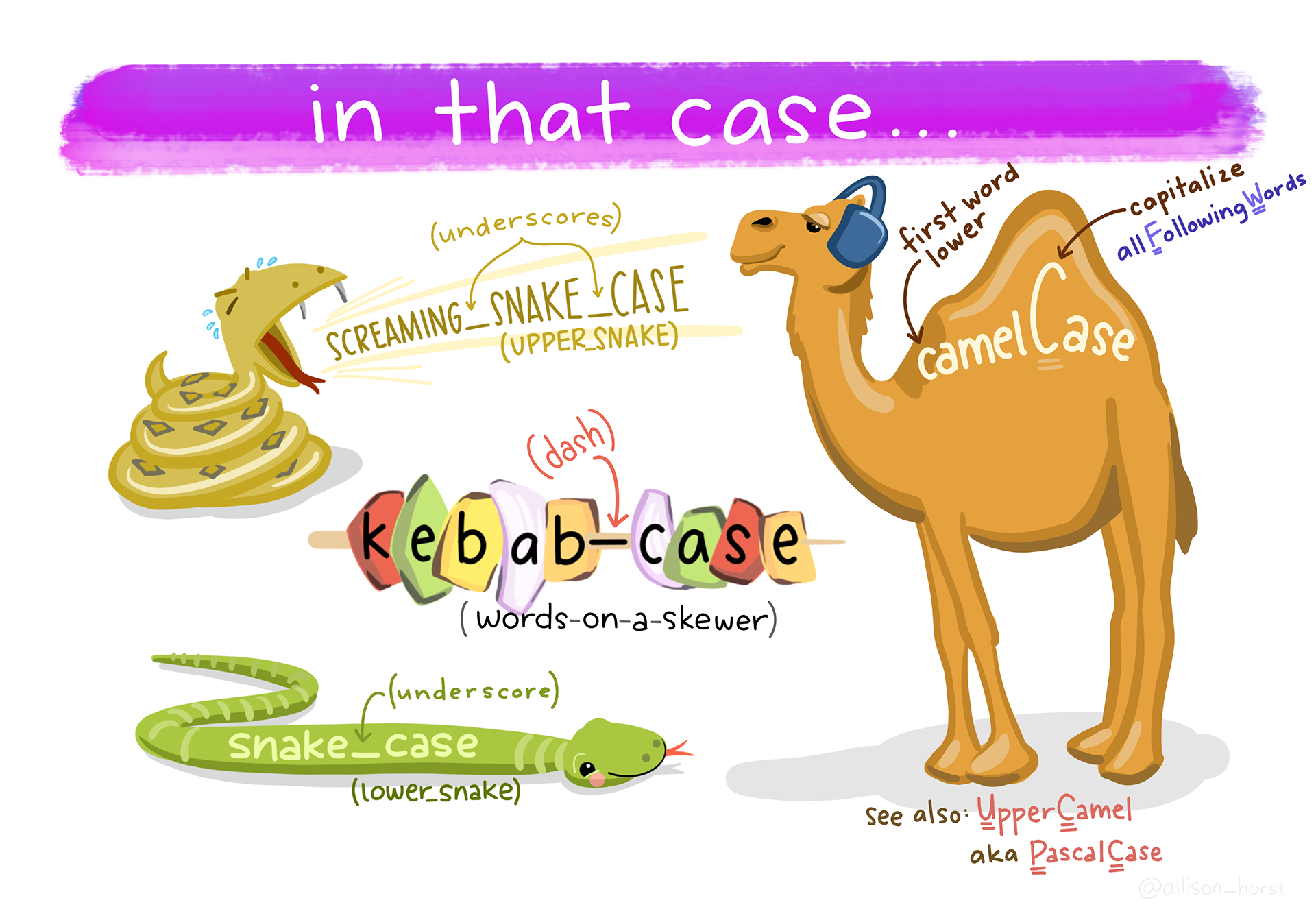
14.2.2 Three common naming styles
You will see these everywhere:
- camelCase (
meanDepth) - PascalCase (
MeanDepth) - snake_case (
mean_depth)
In this course, we will default to snake_case for variables and files. This is strongly recommended for analysis scripts.
14.2.3 General guidelines (the “please don’t make future you angry” list)
These are simple rules that prevent a shocking number of problems:
| Rule | Why it matters | Example of poor form |
|---|---|---|
| Avoid blank spaces | Spaces complicate coding and break formulas, file paths, and scripting | Mean Depth, Site Name, Final Data.csv |
Omit special symbols like ?, $, *, +, #, (, ), -, /, }, {, |, >, <, etc. |
Many special characters are treated as operators or commands in R, so using them in names can break code or make it harder to work with. | mass(g), count#, Y/N?, depth>10 |
Use _ as a separator |
Underscores are safe, readable, and standard | mean-depth, mean.depth, mean depth |
| Do not begin names with numbers | Names starting with numbers are invalid or confusing in R | 100m_depth, 2023_data |
| Make names unique | Non-unique names cause overwriting and ambiguity | data, data2, final, final_final |
| Be consistent with case | R is case-sensitive; inconsistency causes silent bugs | Depth, depth, DEPTH in same project |
| Avoid blank rows in data | Blank rows can be misread as data or missing values | Empty rows between observations in CSV |
| Remove comments from data files | Comments belong in scripts or metadata, not raw data | # measured in June inside a CSV |
| Define and document NA values | Ambiguous missing values lead to incorrect analyses | Mixing NA, 0, -999, . without explanation |
| Use clear, documented date formats | Ambiguous dates are easy to misinterpret | 03/04/21 (is this March 4 or April 3?) |
14.2.4 “Bad → Good” examples for variable names
Your variable names should be: - readable at a glance - easy to type - consistent across files and projects
Here are examples aligned with the tidyverse style guide (Hadley Wickham):
A quick translation rule:
- Use units as suffixes:
depth_cm,mass_g,distance_m(do not use parentheses around units) - Use clear boolean names:
yes_noor better:is_adult,has_nest - Prefer meaning over brevity:
percentile_50beatsp50(unlessp50is a defined term in your field)
14.3 Naming conventions for folders (project structure)
14.3.1 Why folder names matter
A good folder structure makes it hard to lose track of: - raw versus processed data - inputs versus outputs - code versus results - files that you can re-create versus ones you cannot (i.e. that you should never overwrite)
In this course, we care about reproducibility, which means: - raw data is always treated as read-only - processed data is created by scripts, acting on raw data, and never manually edited - results are generated by scripts and never manually edited
Folder naming rules (same spirit as variable naming): - lowercase (snake_case) - no spaces - prefer underscores - descriptive names over clever names
14.4 Conventions for legible and consistent R code
14.4.1 Why code style matters
Code style is not about being fancy. It’s about making your thinking legible.
Good style: - reduces bugs - improves peer review (including lab mates and future you) - makes it easier to modify models without breaking things
14.4.2 “Bad → Good” examples for R code and files
These examples follow the tidyverse style guide (Hadley Wickham):
| Bad | Why was it bad? | Good |
|---|---|---|
Fit models.r |
Contains spaces; inconsistent casing; harder to reference in code | fit_models.R |
########## |
No semantic meaning; comments should describe intent, not fill space | # extract elevation ----- |
findat |
Vague name; gives no clue about content or structure | findat_mat |
findat_mat |
Redundant and unclear; does not describe what the matrix contains | mass_mat |
A few rules we will use repeatedly:
14.4.3 Use nouns for objects and verbs for functions
- Objects (things):
pufferi,pufferi_clean,model_gam - Functions (actions):
clean_data(),fit_model(),plot_effects()
14.4.4 Make “intermediate objects” obvious
If an object is temporary, label it like it is: - *_raw, *_clean, *_long, *_wide, *_summary
14.4.5 Prefer readable code over clever code
14.4.6 A final note about reproducible analyses
In workflow tools like targets, names are not simply cosmetic. They define nodes in a dependency graph (i.e. the ways all your inputs, processes, and outputs are connected). If names are unclear, the graph is unclear. If the graph is unclear, the workflow is not reproducible.
14.5 Your “minimum standard” checklist (use this before you submit anything)
14.6 Practice task (recommended)
- Pick one of your current projects (or a past project).
- Rename:
- 5 confusing variables
- 3 confusing files
- 1 confusing folder
- Make one small style improvement:
- consistent naming
- readable comments
- break one long chunk into 2–3 meaningful steps
The goal is not perfection. The goal is to start practicing now.
